Pulling Back the Curtains on CRISPR Screens
What is a CRISPR screen, and why do scientists love them?
In molecular biology we like to do a lot of screening. Not screening for health problems or a diagnostic test, but we rather we tend to screen libraries looking for something in particular. For pharmacological studies, we can screen libraries containing thousands of small molecules with the potential to be drugs. For CRISPR screens however, libraries are quite different.
CRISPR (clustered regularly interspaced short palindromic repeats) is a gene editing system that was initially found in bacteria. But how did the CRISPR system make its way from bacteria to being a core tool of researchers trying to understand the genetic cause of disease on a large scale?
Today I’m going to discuss the origins behind CRISPR screening, how they work, and how I use them to uncover the biology behind ovarian cancer.
CRISPR-cas systems have been used for gene editing in the lab, and in patients!
I’ve described gene editing in previous posts (which you can read here), but think of CRISPR as a sequence, or set of letters, in our collection of genes which is also referred to as the genome. This sequence allows cas to be a pair of active molecular scissors. Simply puy CRISPR is acting like a power generator to guide the molecular scissors of the cas enzyme to cut DNA at the desired genomic location. There are multiple cas enzymes, with the most popular being cas9, but there are also other like cas12, cas13 etc.
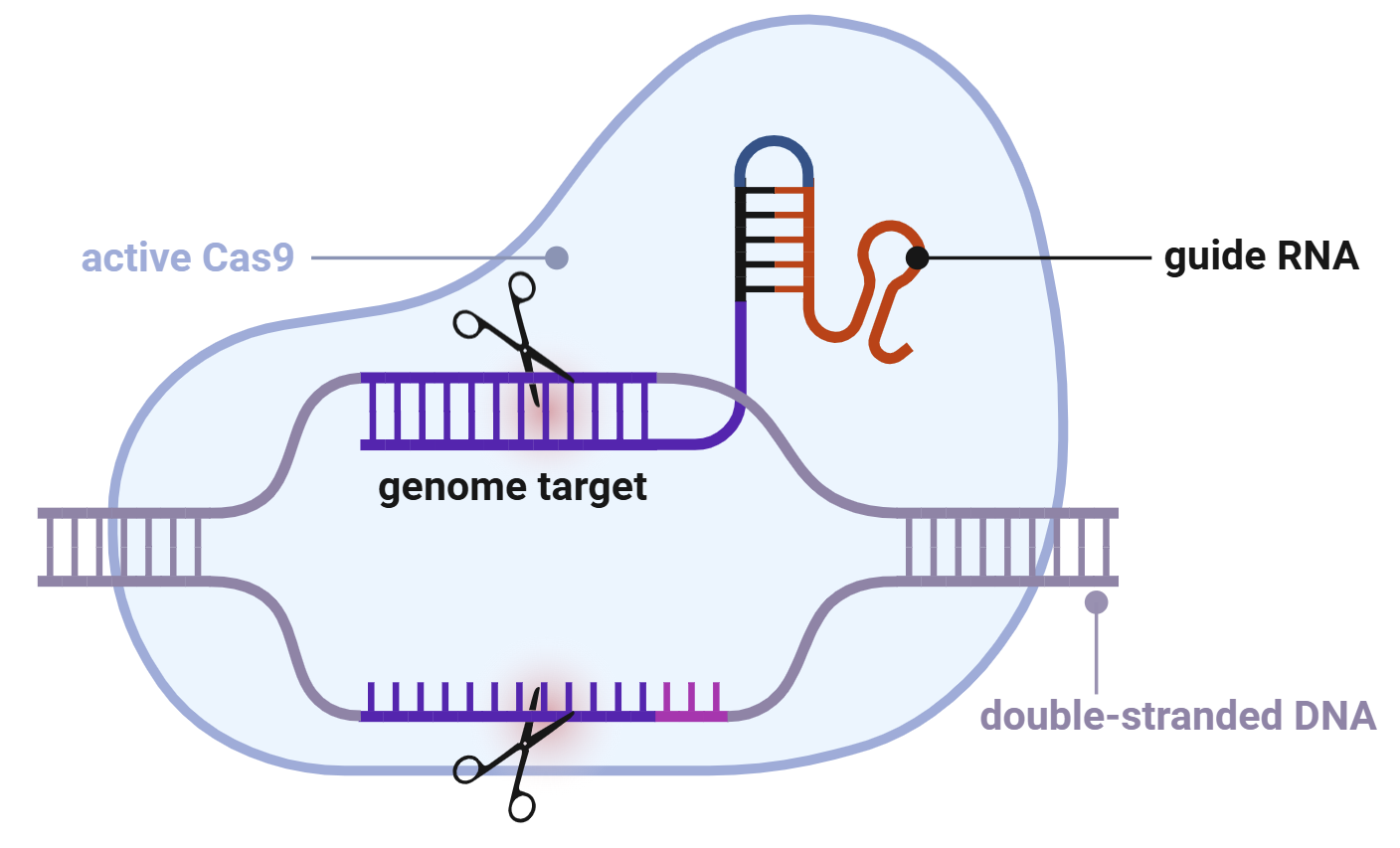
So how does CRISPR-cas9 know where to cut? The system uses single guide RNAs, also referred to as sgRNAs or just guides. The guide RNA guides the cas enzyme to the region of DNA where it will cut. This technology has a lot of awesome spin-offs. First, there are second generations CRISPR-cas enzymes called nickases that cut only one strand of DNA instead of both. These nickases can change single errors in our genomic code, and have even successfully treated a 7 month old boy with a rare metabolic disease last summer.
Other technologies that have modified the original CRISPR-cas9 system are CRISPR activation (or CRISPRa), CRISPR inhibition (CRISPRi), and CRISPR knockout (CRISPR-KO). Think of CRISPRa and CRISPRi like the twin siblings of CRISPR-cas9, while CRISPR-KO is their evil cousin.
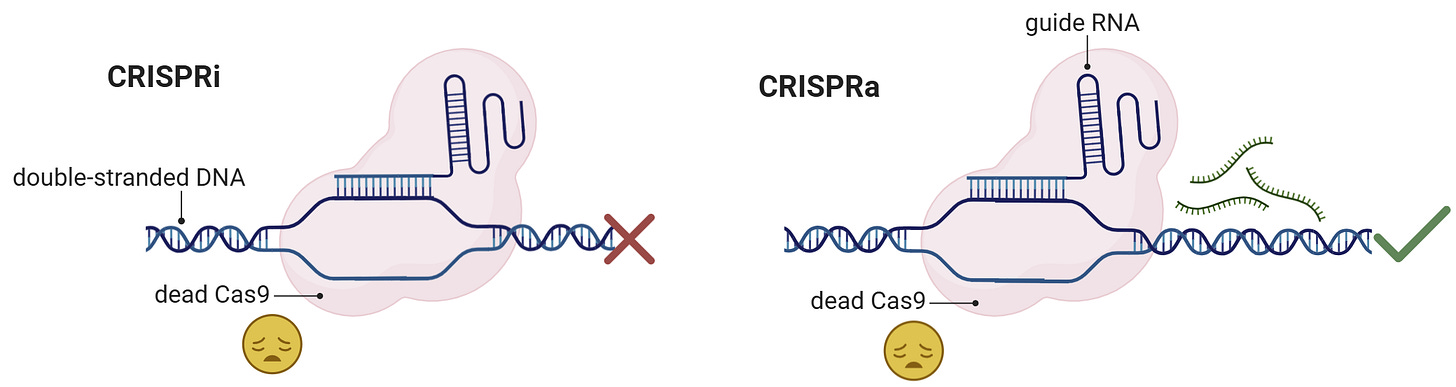
CRISPRa and CRISPRi use a dead cas9 which cannot make cuts in double stranded DNA. This dead-cas9 has special machinery attached to it: for CRISPRa it has activators that increase the targets expression, while CRISPRi has silencers that sit on the target region to silence its expression. CRISPR-KO on the other hand still uses an active cas9 but cuts the desired DNA so that the target gene is not read anymore.
The basics of CRISPR screening:
CRISPR screening allows scientists to interrogate thousands of genes in an unbiased way using the CRISPR cas genetic perturbation system. This experimentation makes up the core of functional genomics research which aims to understand each gene function and their interactions.
Choose any model you like: human brain cells in a dish, mouse models of melanoma, or patient immune cells. Then, pick your CRISPR screening tool of choice – do you want to look at oncogenes or tumour suppressors in cancers? Maybe you want to search for the genetic causes of dementia? Each scientific question requires a different CRISPR variant (i,a, or KO). So now that you’ve picked your research question and model, you need to ask which genes you want to interrogate. To do that you will need to create a library.
What is a library in CRISPR screening?
For CRISPR screens, a library is a collection of guide RNAs that can be customized for every experiment. In practice, they are small tubes of purified RNA that look just like water, with a bit of a higher viscosity. The amazing thing about libraries is that you get to decide what genes to interrogate in your experiment. This can vary from all the genes in the human genome (almost 20 000!!!) or maybe a subset of a few hundred or thousand genes that are involved in your disease of interest. Using bioinformatic tools to probe patient data, you can ensure your experiment is accurate and reflects clinical cases.
CRISPR screens work by changing one gene’s activity in only one cell at a time
We know that cas enzymes are like scissors, cutting DNA strands and knocking out a gene. With dead cas, you can activate or inhibit a gene of interest. Once you complete your screen, you send your sample for deep sequencing, which reads all the guide RNAs present in your sample.
In mouse models of cancer that I’ve worked with, instead of collecting cells, we collect tumours that grow after our guide libraries have been added. The deep sequencing results tell us which guide RNAs were present in the tumour, and thus which genes from our collection of interrogated genes caused cancer in the mice.
How do I use CRISPR-cas screening to study ovarian cancer?
By using cancer patient data, I can identify patterns of genes that are usually overexpressed and repressed in cancer patients. Having the ability to identify these genes is great, but we don’t know which ones are actually causing cancer and which ones are just along for the ride, or bystanders.
By injecting the guide libraries into a mouse model of ovarian cancer with cas9, I can find out which genes might cause or repress tumour growth in a mouse model. For example, say after deep sequencing of a CRISPRi mouse tumour, I see that the gene CHEK2 (involved in DNA repair) was the guide from that cell population. Since tumours grew when CHEK2 was inhibited, we can infer that it usually acts as a tumour suppressor, and inhibiting the gene in a mouse caused cancer to grow. And the opposite would be true for a guide found from a CRISPRa tumour. We would probably hypothesize its identity as an oncogene.
CRISPR screening is a powerful unbiased approach to understand gene function
I hope I’ve shown you the power of the CRISPR-cas system for scientific questions and experiments. It’s an amazing tool that allows you to interrogate thousands of genes in almost any model and discover genetic drivers of disease, and it’s my main method of choice for my thesis work.
See you in the next post!


Adele, thanks for making this easy for a non-scientist. Do most labs you know use the different variants of CRISPR? How widely used it, do you know? It sounds like it has so much potential